Research
Interests
I have a broad interest in methods and applications in spatial statistics. In addition to working with spatial point processes (often surrounding intensity estimation), I also have an emerging interest in spatial regressions, including adapting and extending Gaussian conditional autoregressive (CAR) models and similar models for geostatistical data. My areas of research include:
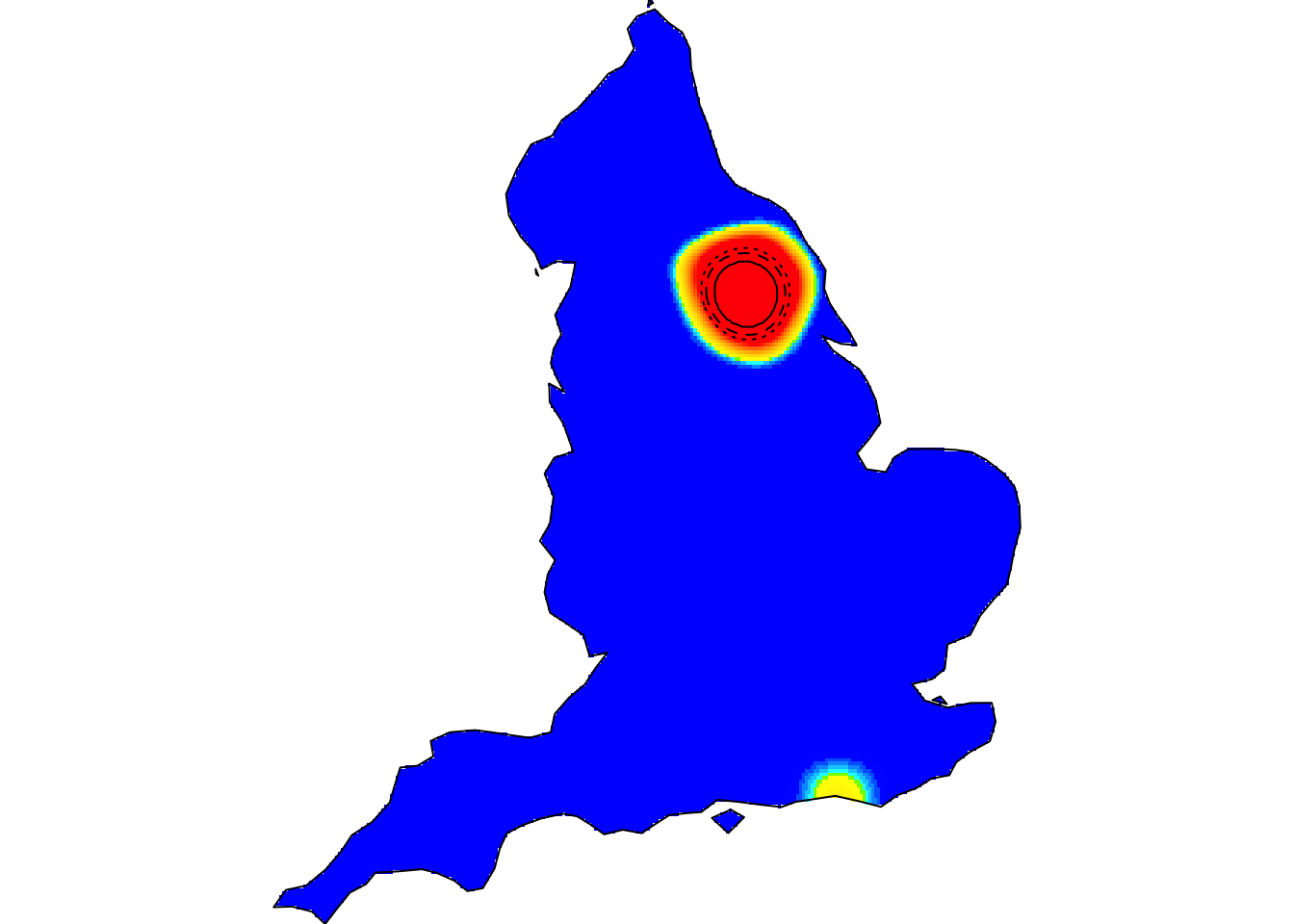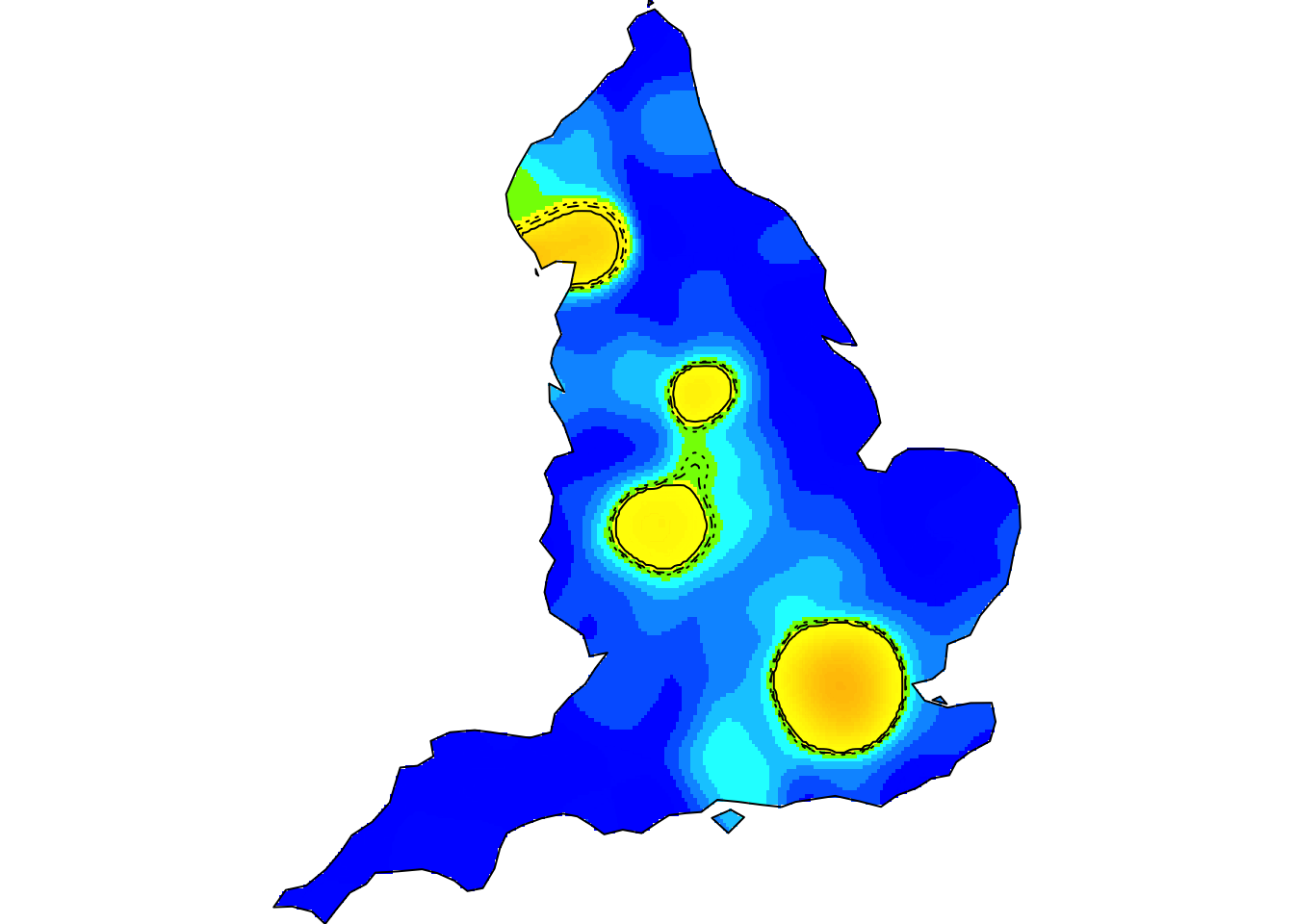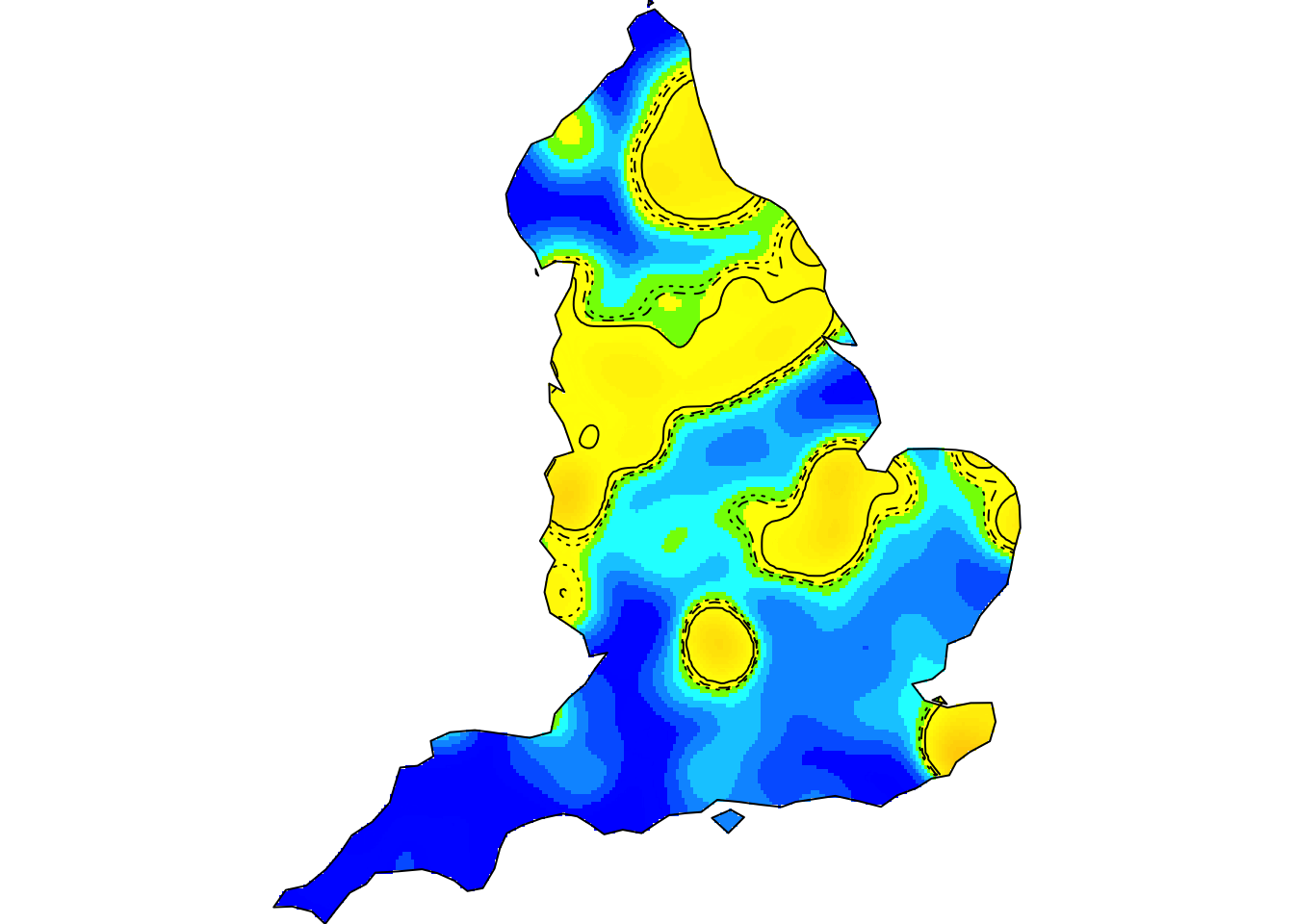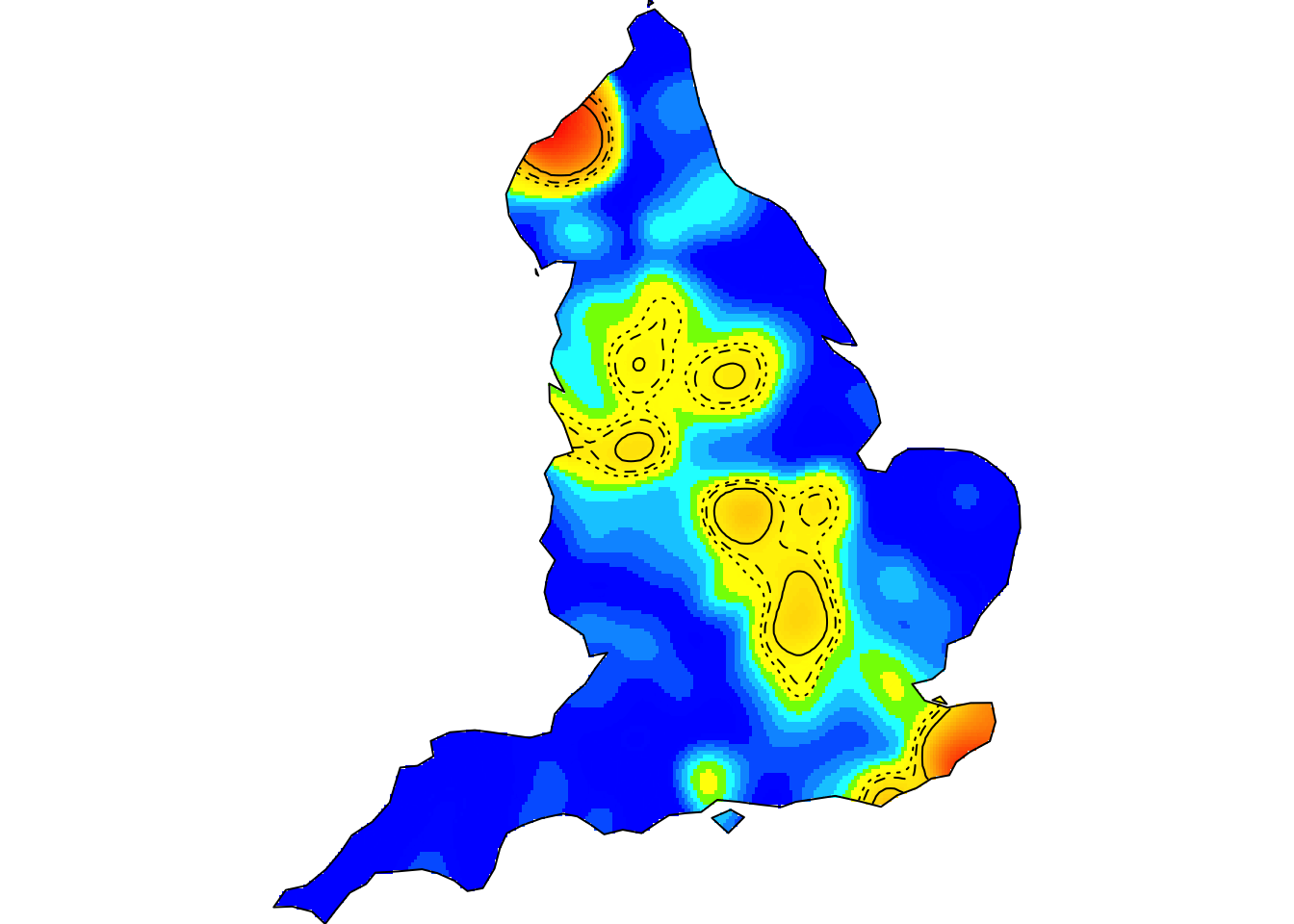
Kernel smoothing, nonparametric intensity estimation (the animation above shows the spatiotemporal relative risk of COVID-19 during the first six months of England’s 2020 outbreak – see refs. [20,27] below)
Spatial point processes and related methodology
Spatial regression / geostatistics
Applications of the above in various fields, such as geographical epidemiology, physiology, geology and ecology
If you have a strong academic record and are interested in working with me as a postgraduate student on topics related to the above, get in touch.
Grants and awards
2024: Littlejohn Research Award, New Zealand Statistical Association (NZSA).
2023: Principal Investigator, Marsden Fund (Royal Society of New Zealand) Grant 23-UOO-148. “Principled inference for spatial point processes: a unified toolkit”. Associate Investigators Dr Charlotte Jones-Todd (University of Auckland, NZ); Prof Martin Hazelton (University of Otago, NZ); Prof Adrian Baddeley (Curtin University, Australia); Assoc. Prof. Ege Rubak (Aalborg University, Denmark). NZ$818,800; March 2024 - March 2027.
2019: Principal Investigator, Marsden Fund (Royal Society of New Zealand) Grant 19-UOO-191. “A new generation of statistical models for spatial point process data”. Associate Investigators Prof Adrian Baddeley (Curtin University, Australia) and Prof Martin Hazelton (University of Otago, NZ). NZ$810,750; March 2020 - March 2023.
2017: Early Career Award for Distinction in Research, University of Otago.
2015: Principal Investigator, Marsden Fund (Royal Society of New Zealand) Fast-start Grant 15-UOO-092. “Smoothing and inference for point process data with applications to epidemiology”. Associate Investigators Dr Ben Taylor (Lancaster University, UK) and Prof Martin Hazelton (University of Otago, NZ). NZ$345,000; March 2016 - March 2019.
2014: Worsley Early Career Research Award, New Zealand Statistical Association (NZSA).
2012: Principal Investigator, University of Otago Research Grant-in-aid: “Statistical Methods for Spatial Intensity Estimation and their Performance in Epidemiology”. NZ$5,700; Jan 2013 - Dec 2013.
Journal Publications
[41] Davies TM (2026) Gated spatial hidden Markov models for classification in histological samples, with application to muscle fibre-typing, Submitted for publication.
[40] Macdonald BJ, Davies TM, Hazelton ML, Baddeley A (2026) A unified methodology for testing in clustered spatial processes, Submitted for publication.
[39] Zamorano D, Matthaei CD, Meier CI, Davies TM, Ingram T, Romero U (2026) Estimating dispersal kernels for stream periphyton using drifting lake algae, Submitted for publication.
[38] Davies TM (2026) Editorial: 25 YeaRs (Special Issue), Australian and New Zealand Journal of Statistics [in press].
[37] Baddeley A, Davies TM, Hazelton ML (2025) An improved estimator of the pair correlation function of a spatial point process, Biometrika 112 2 asaf021.
[36] Macdonald BJ, Davies TM, Hazelton ML (2025) Bandwidth selection for kernel intensity estimators for spatial point processes, Scandinavian Journal of Statistics 52 3 1111-1137.
[35] Macdonald BJ, Davies TM, Hazelton ML (2023) Feasibility of Monte-Carlo maximum likelihood for fitting spatial log-Gaussian Cox processes, Spatial Statistics 56 100759.
[34] Redmond AK, Davies TM, Schofield MR, Sheard PW (2023) New tools for investigation of muscle fiber-type spatial distributions across histological sections, Skeletal Muscle 13 7 1-11.
[33] Baddeley A, Davies TM, Hazelton ML, Rakshit S, Turner R (2022) Fundamental problems in fitting spatial cluster process models, Spatial Statistics 52 100709.
[32] Baddeley A, Davies TM, Rakshit S, Nair G, McSwiggan G (2022) Diffusion smoothing for spatial point patterns, Statistical Science 37 1 123-142.
[31] Crump JA, Davies TM (2022) Towards equitable scheduling of global health teleconferences: a spatial exploration of the world’s population and health by time zone, BMJ Open 12 e056696.
[30] Davies TM, Banerjee S, Martin AP, Turnbull RE (2022) A nearest-neighbour Gaussian process spatial factor model for censored, multi-depth geochemical data, Journal of the Royal Statistical Society Series C (Applied Statistics) 71 4 1014-1043.
[29] Hazelton ML, Davies TM (2022) Pointwise comparison of two multivariate density functions, Scandinavian Journal of Statistics 49 4 1791-1810.
[28] Baddeley A, Nair G, Rakshit S, McSwiggan G, Davies TM (2021) Analysing point patterns on networks – a review, Spatial Statistics 42 100435.
[27] Elson R, Davies TM, Lake IR, Vivancos R, Blomquist PB, Charlett A, Dabrera G (2021) The spatio-temporal distribution of COVID-19 infection in England between January and June 2020, Epidemiology and Infection 149 e73 1-6.
[26] Elson R, Davies TM, Jenkins C, Vivancos R, O’Brien SJ, Lake IR (2020) Application of kernel smoothing to estimate the spatio-temporal variation in risk of STEC O157 in England, Spatial and Spatio-temporal Epidemiology 32 100305.
[25] Davies TM, Lawson AB (2019) An evaluation of likelihood-based bandwidth selectors for spatial and spatiotemporal kernel estimates, Journal of Statistical Computation and Simulation 89 7 1131-1152.
[24] Davies TM, Schofield MR, Cornwall J, Sheard PW (2019) Modelling multilevel spatial behaviour in binary-mark muscle fibre configurations, Annals of Applied Statistics 13 3 1329-1347.
[23] Rakshit S, Davies TM, Moradi MM, McSwiggan G, Nair G, Mateu J, Baddeley A (2019) Fast kernel smoothing of point patterns on a large network using 2D convolution, International Statistical Review 87 3 531-556.
[22] Davies TM, Baddeley A (2018) Fast computation of spatially adaptive kernel estimates, Statistics and Computing 28 4 937-956.
[21] Davies TM, Flynn CR, Hazelton ML (2018) On the utility of asymptotic bandwidth selectors for spatially adaptive kernel density estimation, Statistics & Probability Letters 138 75-81.
[20] Davies TM, Marshall JC, Hazelton ML (2018) Tutorial on kernel estimation of continuous spatial and spatiotemporal relative risk, Statistics in Medicine 37 7 1191-1221.
[19] Davies TM, Jones K, Hazelton ML (2016) Symmetric adaptive smoothing regimens for estimation of the spatial relative risk function, Computational Statistics & Data Analysis 101 12-28.
[18] Davies TM, Sheard PW, Cornwall J (2016) Letter to the Editor: Comment on Makino et al. and observations on spatial modeling, Anatomical Science International 91 4 423-424.
[17] Farrell S, Davies TM, Cornwall J (2015) Use of clinical anatomy resources by musculoskeletal outpatient physiotherapists in Australian public hospitals: A cross-sectional study, Physiotherapy Canada 67 3 273-279.
[16] Fletcher JGR, Stringer MD, Briggs CA, Davies TM, Woodley SJ (2015) CT morphometry of adult thoracic intervertebral discs, European Spine Journal 24 10 2321-2329.
[15] Smith BA, Davies TM, Higham CFW (2015) Spatial and social variables in the Bronze Age phase 4 cemetery of Ban Non Wat, Northeast Thailand, Journal of Archaeological Science: Reports 4 34 362-370.
[14] Taylor BM, Davies TM, Rowlingson BS, Diggle PJ (2015) Bayesian inference and data augmentation schemes for spatial, spatiotemporal and multivariate log-Gaussian Cox processes in R, Journal of Statistical Software 63 7 1-48.
[13] Cornwall J, Davies TM, Lees D (2013) Student injuries in the dissecting room, Anatomical Sciences Education 6 6 404-409.
[12] Davies TM (2013) Jointly optimal bandwidth selection for the planar kernel-smoothed density-ratio, Spatial and Spatio-temporal Epidemiology 5 1 51-65.
[11] Davies TM (2013) Scaling oversmoothing factors for kernel estimation of spatial relative risk, Epidemiological Methods 2 1 67-83.
[10] Davies TM, Bryant DJ (2013) On circulant embedding for Gaussian random fields in R, Journal of Statistical Software 55 9 1-21.
[9] Davies TM, Cornwall J, Sheard PW (2013) Modelling dichotomously marked muscle fibre configurations, Statistics in Medicine 32 24 4240-4258.
[8] Davies TM, Hazelton ML (2013) Assessing minimum contrast parameter estimation for spatial and spatiotemporal log-Gaussian Cox processes, Statistica Neerlandica 67 4 355-389.
[7] Taylor BM, Davies TM, Rowlingson BS, Diggle PJ (2013) lgcp - An R package for inference with spatial and spatiotemporal log-Gaussian Cox processes, Journal of Statistical Software 52 4 1-40.
[6] Zhang ZJ, Davies TM, Gao J, Wang Z, Jiang QW (2013) Identification of high-risk regions for schistosomiasis in the Guichi region of China: an adaptive kernel density estimation-based approach, Parasitology 140 7 868-875.
[5] Zhang ZJ, Chen DM, Chen Y, Davies TM, Vaillancourt JP, Liu WB (2012) Risk signals of an influenza pandemic caused by highly pathogenic avian influenza subtype H5N1: Spatio-temporal perspectives, Veterinary Journal 192 3 417-421.
[4] Davies TM, Hazelton ML, Marshall JC (2011) sparr: Analyzing spatial relative risk using fixed and adaptive kernel density estimation in R, Journal of Statistical Software 39 1 1-14.
[3] Sanson RL, Harvey N, Garner MG, Stevenson MA, Davies TM, Hazelton ML, O’Connor J, Dubé C, Forde-Folle KN, Owen K (2011) Foot-and-mouth disease model verification and ‘relative validation’ through a formal model comparison, OIE Scientific and Technical Review 30 2 527-540.
[2] Davies TM, Hazelton ML (2010) Adaptive kernel estimation of spatial relative risk, Statistics in Medicine 29 23 2423-2437.
[1] Hazelton ML, Davies TM (2009) Inference based on kernel estimates of the relative risk function in geographical epidemiology, Biometrical Journal 51 1 98-109.